RNA-SEQ
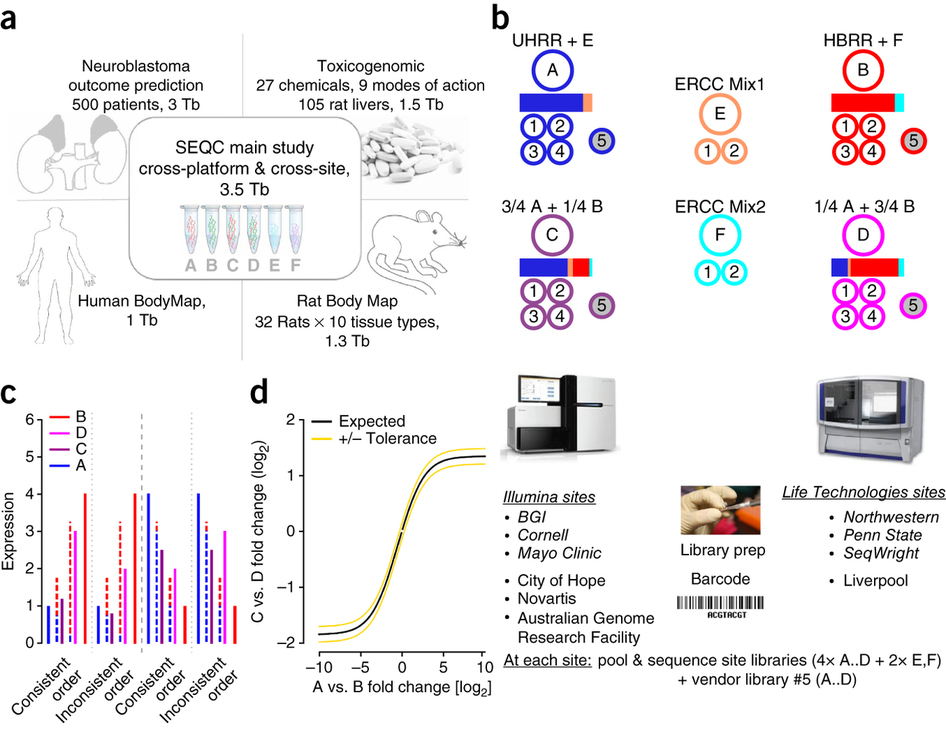
We are happy to post here the results, methods, data, scripts, and phalanx of papers for the Food and Drug Administration’s (FDA) Sequencing Quality Control (SEQC) Study on RNA-Sequencing (RNA-Seq) and the Association of Biomolecular Resource Facilities (ABRF) Phase I Study on Next-Generation Sequencing (NGS), which focused on RNA-seq. These 10 complementary studies have focused on all aspects of RNA-sequencing, including: RNA quality, library preparation, sequencing, alignment, quantification, normalization, integration, and concordance. The overarching goal was to establish robust measures and methods for of inter-site and inter-platform RNA-seq concordance, which is critical for clinical applications, self-evaluation/improvement, and technology and informatics development.